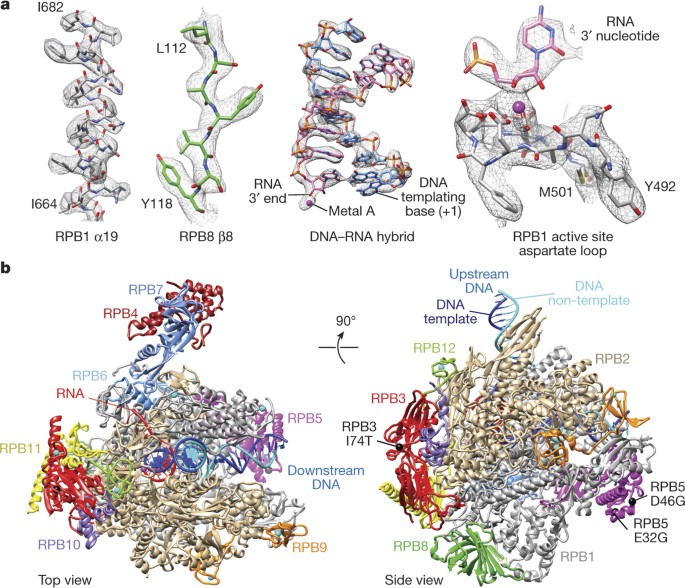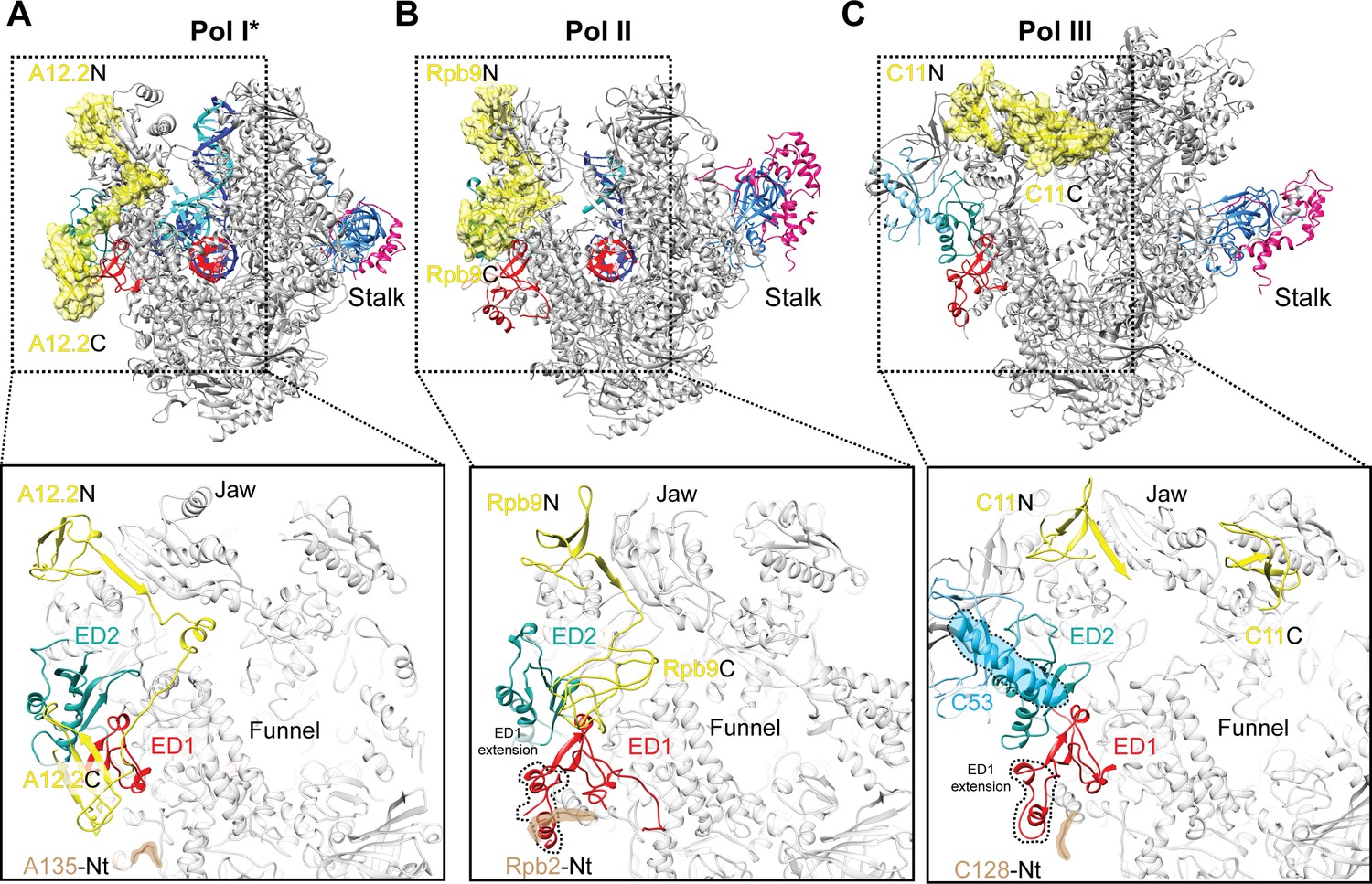
The cryo-EM structure of a 12-subunit variant of RNA polymerase I reveals dissociation of the A49-A34.5 heterodimer and rearrangement of subunit A12.2 | eLife
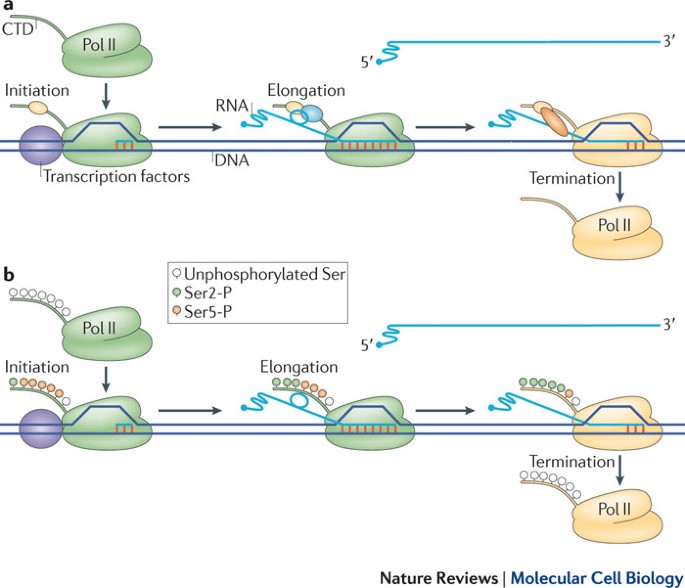
Unravelling the means to an end: RNA polymerase II transcription termination | Nature Reviews Molecular Cell Biology
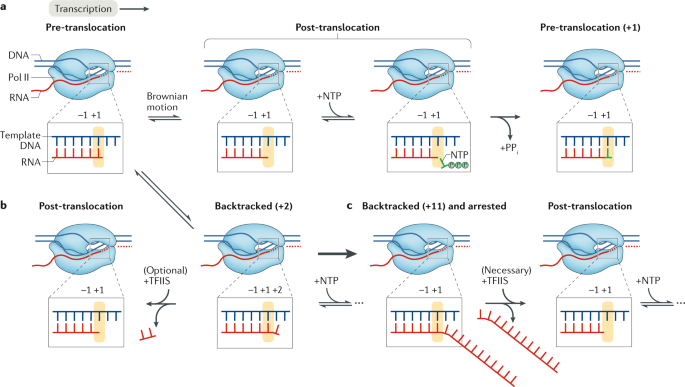
Causes and consequences of RNA polymerase II stalling during transcript elongation | Nature Reviews Molecular Cell Biology

Phosphorylation-Regulated Binding of RNA Polymerase II to Fibrous Polymers of Low-Complexity Domains: Cell

RNA Pol II Length and Disorder Enable Cooperative Scaling of Transcriptional Bursting - ScienceDirect

Control of RNA Pol II Speed by PNUTS-PP1 and Spt5 Dephosphorylation Facilitates Termination by a “Sitting Duck Torpedo” Mechanism - ScienceDirect

Schematic representation of the RNA polymerase II holoenzyme bound to a... | Download Scientific Diagram

RNA Polymerase II Phosphorylated on CTD Serine 5 Interacts with the Spliceosome during Co-transcriptional Splicing - ScienceDirect

Structure and function of the pol II transcription machinery | Taatjes Lab | University of Colorado Boulder

Cell- and Polymerase-Selective Metabolic Labeling of Cellular RNA with 2′-Azidocytidine | Journal of the American Chemical Society

RNA polymerase II clusters form in line with surface condensation on regulatory chromatin | Molecular Systems Biology